rna-tools.online
diffpdb
files from another job e.g., 614e0a1b
The raw output files for each step of the pipeline can be found here and downloaded as a ZIP file here.
Documentation
The tool uses 'diff' comment for text-based comparison of files. Read more on 'diff' here
diffpdb checks the consistency between annotations of two structural files. The tool ignores 3D coordinates of atoms and compares only text-content of two files in the PDB to identify the difference in the annotation of atoms, missing atoms (missing the O2′ atom) and missing fragment (shown on the left side with the gray-red bar).
ATOM 8 C2' G A 57 ATOM 8 C2' G A 57
# this is when these two files differe, this is the place when O2' atom exists on the left
# (crystal structure) and not on the right (SimRNA model)
ATOM 9 C1' G A 57 | ATOM 9 O2' G A 57
ATOM 10 N1 G A 57 | ATOM 10 C1' G A 57
diffpdb can be also used locally with a diff viewer of choince. Here the detection of missing O2' atom in one of the structure can be easily detected. This is
Read more on this puzzle here https://github.com/mmagnus/RNA-Puzzles-Standardized-Submissions/tree/master/rp17
Slightly different presentation of the same information can be viewed with other diff viewers, like DiffMerge here: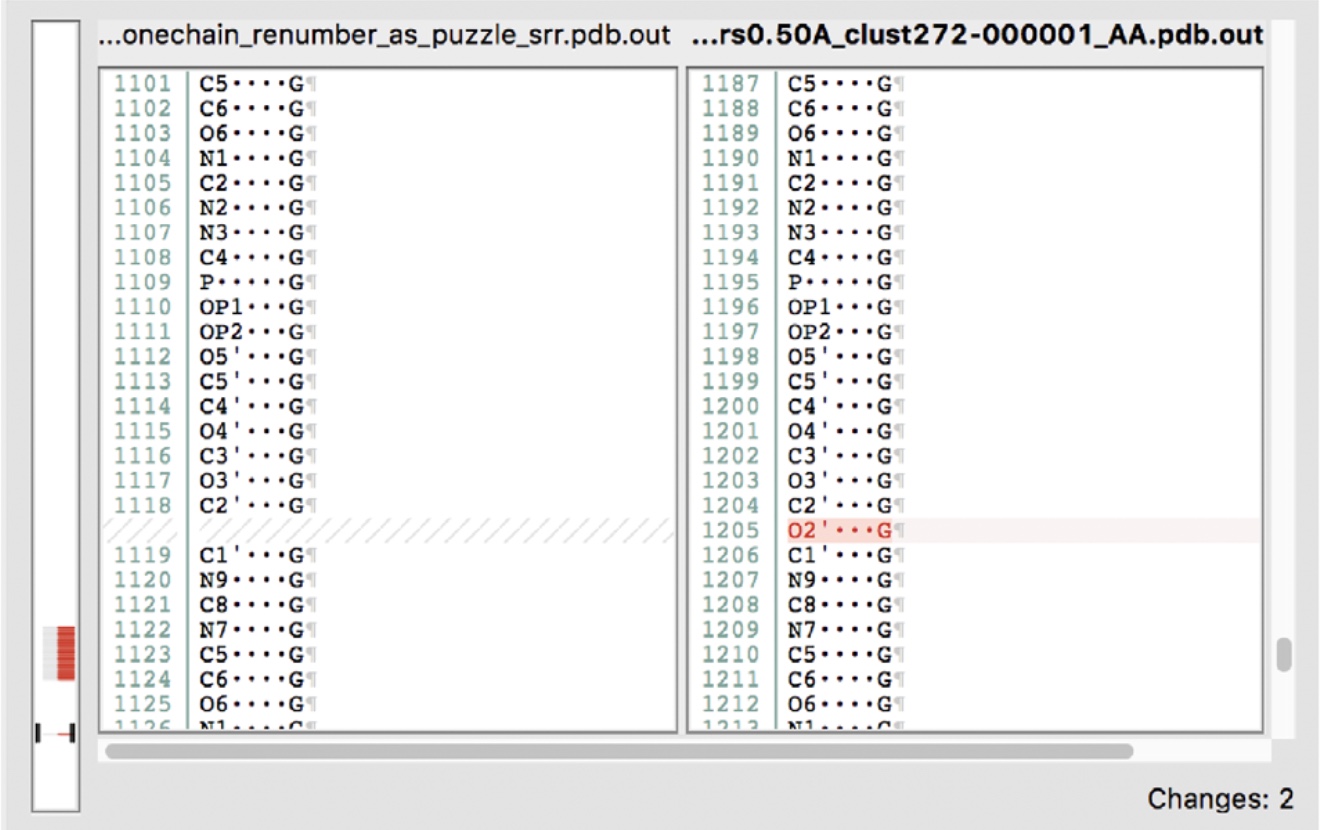
We are exploring using https://piwik.pro to track usage of the server for grant proposals, scientific presentations. Please write to us if there is any problem with your job mail us [behind Apple Hide My Mail].
